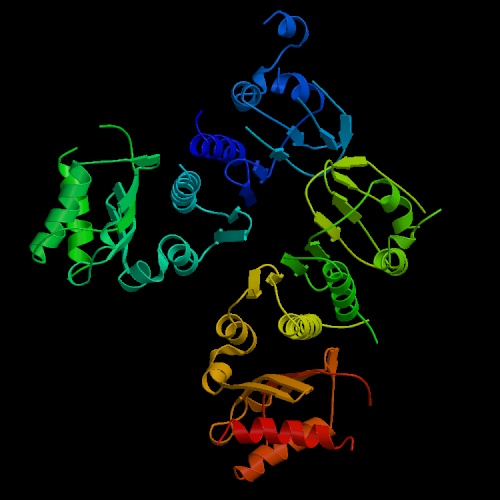
(a) ID number : 1BKN
(b) Name of the molecule : Crystal Structure Of An N-Terminal 40Kd Fragment Of E. Coli DNA Mismatch Repair Protein Mutl
(c) Experimental method for structure determination : X-ray Diffraction
(d) Resolution of the structure : 2.90A
(a) number of protein chains : 2 ( A and B)
(b) number of helix in one protein chain : A chain : 11 helix , B chain : 10 helix
(c) the first and last residues in the helix 6 : first : T and last : A
Class : Alpha and beta proteins (a+b) Mainly antiparallel beta sheets (segregated alpha and beta regions)
Fold :
ATPase domain of HSP90 chaperone/DNA topoisomerase II/histidine kinase
8-stranded mixed beta-sheet;
2 layers: alpha/beta
Superfamily : ATPase domain of HSP90 chaperone/DNA topoisomerase II/histidine kinase
Family
: DNA gyrase/MutL, N-terminal
domain

Assignment
7
Protein Structure Database -PDB-.
- To do
Protein Structure data search
- To understand protein structure data file
1. Search human Mlh1 or its homologous proteins in Protein Data Bank (PDB) database.
You just need to search for protein structure, do not find complex structure.
Give its (a) ID number, (b) Name of the molecule, (c) experimental method for
structure determination and (d) Resolution of the structure.
Ans :
2. Show its structure in ribbons form (400x400).
3. Give the structural informations
of this PDB file (a) number of protein chains, (b) number of helix in one protein
chain, (c) the first and last residues in the helix 6 .
Ans :
4. Give the Class, Fold, Superfamily
and Family of this protein in Structural Classification of Proteins (SCOP).
Ans :
5. (a) Show Ramachandran plot for
this PDB file. (b) How many residues are in the most favoured regions of Ramachandran
plot?
Ans :